Food Safety
Evolving pathogen testing methods offer better, faster alternatives
New technologies offer more speed, better accuracy and ease of use for pathogen testing.

The CDC’s Dr. Andrew Huang performs research involving metagenomics in his lab. Source: CDC.
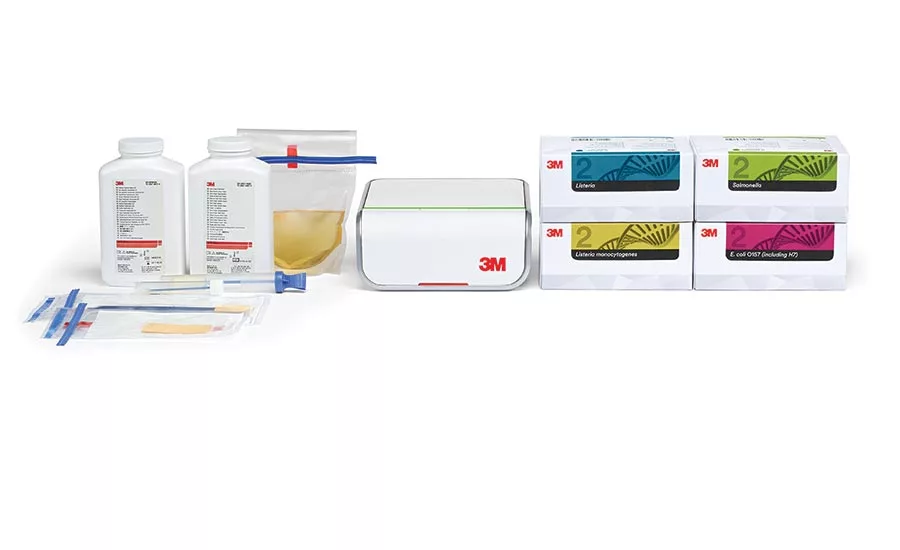

3M’s family of pathogen testing products, used for screening samples taken from both finished products and from processing environments. Source: 3M.
Clear Labs set out to build a software platform and analytics platform powered by next-gen DNA testing for food retailers and manufacturers. Source: Clear Labs.
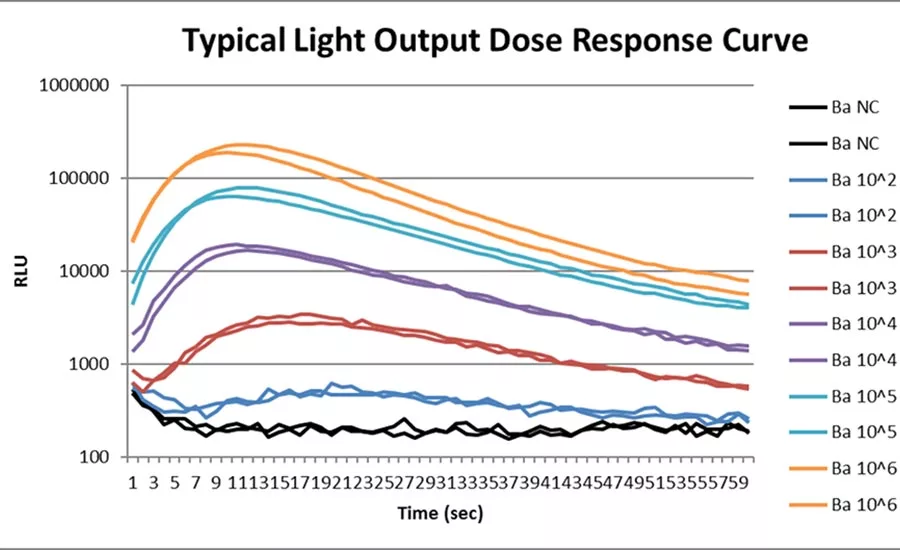
This graph shows the typical light output dose response curve for PathSensors CANARY technology using the Zephyr (powder/solution samples) system. This graph is for bacteria at different concentrations. As observed from the x-axis, this light detection is happening during a one-minute timespan and is able to detect down to 10^2 concentration levels of bacteria. Source: PathSensors.

This graphic shows how advanced molecular detection has allowed scientists to detect more clusters of illness, solve outbreaks faster with fewer cases and link cases to unlikely sources. Source: CDC.
Silent and undetectable to the naked eye, pathogens are the nemeses of processors. If a test result comes in positive, it can mean, at the very least, a bumpy road ahead for an uncertain amount of time. A lot is riding on these food safety tests, and more of them are being conducted than, perhaps, ever before, thanks, in part, to the Food Safety Modernization Act’s emphasis on preventative controls.
In an interview conducted after Sabra Dipping Company issued a voluntary recall in November 2016 due to a positive Listeria monocytogenes test result in its factory, but not found in the finished product, company officials made it clear that hummus is not particularly susceptible to foodborne pathogens.
Article Index:
- Traditional foodborne pathogen detection methods
- Rapid pathogen detection systems
- Whole genome sequencing
- The benefits of rapid detection methods
- Sidebar: Rapid detection of bacterial outbreaks
“I think it’s a reflection of the increased scrutiny of FSMA, with FDA trying to solve problematic food cases that so far haven’t been solved,” says Tulin Tuzel, chief technology officer for Sabra. “And the increased scrutiny is a good thing, but I wouldn’t say hummus is more susceptible to Listeria. In fact, it might be a little less susceptible since it’s an intermediate pH food; it’s not a high pH food that can grow bacteria easily.”
In response to the result, Sabra vowed to continue employing all the food safety processes already in place, such as testing incoming ingredients and environmental testing, as well as testing finished product and holding the product until all is confirmed safe. Along with its in-house food safety experts, Sabra also brought in third-party advisors to get opinions of what additional food safety processes could be done or updated.
“There are always new technologies coming out and new things you can learn, and so we immediately put in place the new measures,” says Ilya Welfeld, who handles media relations for Sabra. “And on an ongoing basis, those measures will continue to be evaluated.”
One area of technology development is the actual tests. There is a wide array of diagnostic platforms available to processors today to help obtain a fast and reliable food safety testing program. Also, new technology being released shows a promising future for the speed and accuracy of pathogen testing, offering more ways to integrate automated solutions into preventative controls.
Traditional foodborne pathogen detection methods
In the food safety testing market, there are a number of diagnostics that processors can run in addition to finding foodborne pathogens, such as testing DNA for GMOs, verifying ingredient authenticity and screening for allergens. However, the most performed of these is pathogen detection.
The current status of the food safety testing market is largely still dominated by sending off culture-based samples to a lab. This can pose a problem for processors, even if this testing process method is entrenched at a facility and has been used for many years. The results can take days for microbiological approval.
“When you’re doing your testing with a lab, while you’re waiting on the results, the question really is, what behavior do you exhibit?” says Michael Koeris, co-creator of Sample6, a rapid pathogen diagnostic system. “Do you ship at risk, or do you test and hold? Both aren’t optimal, but you don’t have a choice.”
That is, he says, processors don’t have any other choices if they are waiting on the lab results with traditional pathogen detection methods. But if the results are available within the same shift, then a company can declare with confidence that there is a problem or there is not.
“The higher value that rapid testing delivers is to quickly get the result and avoid days of holding and operational costs,” he says.
A worse scenario yet is if a processor decides to ship without getting the results first and then finds a positive. In this case, not only do you have all the initial transportation costs, you also have the recall costs, which can be very expensive.
“Not only that, your product contamination insurance, PCI, can kick in,” Koeris says.
PCI is the insurance that manages the losses that are incurred by recalling and replacing your product. Koeris says it can be the replacement of the food product that can be more challenging since retailers’ contracts usually specify that processors have to supply them with product at the processor’s cost.
“Secondly, and more importantly, you may make people sick. That is a risk that most people don’t want to think about, but it is real,” Koeris says. “If you have Listeria in your environment, or it’s in your product and you’re shipping at risk, you may kill people. That’s your brand and your livelihood, and it could possibly mean jail time.”
Recently, a ConAgra Brands subsidiary agreed to pay $11.2 million to resolve a criminal investigation into a multi-state Salmonella outbreak blamed on contaminated peanut butter. That kind of fine can be fatal to a small to mid-sized company and illustrates, as Koeris says, that “your brand risk is very real.”
Rapid pathogen detection systems
In the 1970s, research on immunoassays paved the way for the commercial availability of enzyme-linked immunosorbent assay (ELISA) technology. This rapid pathogen detection test is based on the nature of antigen/antibody binding and involves placing food or environmental samples in a solution and observing for an “antibody-antigen-antibody sandwich.” Most available rapid detection assays provide results in less than 24 hours.
“Time-to-result continues to be the arms race among pathogen detection technology providers,” says Adriana Robayo, pathogens global marketing manager for 3M. “Most testing equipment manufacturers are focused on making speed advances within the enrichment stage, as the process of growing bugs to detectable levels often takes 18-24 hours versus the comparatively shorter times needed to actually prepare, run and analyze the tests.”
For instance, Sample6 has developed an enrichment-free pathogen diagnostic system. The Bioillumination platform uses synthetic biology to target bacteria. So far, the testing is approved for detecting Listeria and is in the process of being certified for the detection of Salmonella and E. coli by the end of 2017. The total time-to-result for Sample6 testing is six to eight hours.
PathSensors, a biotechnology company focused on environmental and foodborne disease testing, has developed a variety of pathogen detection platforms using technology developed at MIT Lincoln Labs. When the platform, specifically its Zephyr system, is paired with the company’s biosensors, the solution provides rapid screening for foodborne pathogens, including Listeria, Salmonella, Campylobacter and others.
The method uses immunoassays, like ELISA, to detect pathogens, but has eliminated a number of steps, like the sandwiching antibodies process, by using CANARY (Cellular Analysis and Notification of Antigen Risks and Yields), a cell-based biosensor technology.
“[In the] PathSensors assay process, these things are happening naturally—and nearly instantaneously—within the cell,” says Jack Fritzinger, office systems manager for PathSensors.
CANARY incorporates pathogen-specific antibodies expressed on the biosensor surface triggering, in the presence of a pathogen, an intracellular calcium release that in turn activates bioluminescent proteins. The company’s assays have been designed to be much more sensitive, which shortens enrichment time; the CANARY technology produces average time-to-result in less than five minutes.
“And this speed can be maintained without sacrificing our 100-1,000 FU limit of detection,” he says.
Shown in GRAPH 1, the CANARY technology using the Zephyr system detects bacteria down to 10^2 concentration levels during a one-minute timespan.
“This level of specificity in an assay is rare and unheard of to do within such a small amount of time,” Fritzinger adds.
In the 1980s, research involving polymerase chain reaction (PCR) technology produced the commercial availability of molecular-based detection testing. PCR methods use instrumentation involving temperature cycling to denature a sample’s DNA and detect amplified target DNA through fluorescence, which Robayo says can pose maintenance challenges. So, the 3M Molecular Detection System was created to be a better test.
“3M’s technology contains a unique, high-fidelity DNA polymerase that uses isothermal DNA amplification—amplification at a constant temperature,” she says. “It also targets genes using many more primers than PCR can and employs a special enzyme to help read amplified DNA results in a way that’s continuous and without interruption.”
Combining these two technologies—isothermal DNA amplification and bioluminescence detection—allows them to occur together, providing real-time results in a short period of time. She also says the system is more tolerant of inhibitors in samples and less susceptible to repeat tests.
Whole genome sequencing
Another area of food safety testing that has recently been growing is whole genome sequencing (WGS). Clear Labs, started in 2014 by Sasan Amini and Mahni Ghorashi, has developed the first food industry-specific WGS test based on next-generation sequencing (NGS).
Designed for retailers and manufacturers, Clear Labs built both a software and analytics platform powered by NGS technology to identify specific pathogen strains, link them to similar species and monitor for pathogen-related problems in the food supply.
“We’re helping retailers and manufacturers to build more airtight food safety and quality pipelines,” Ghorashi says. “Whole genome sequencing is being used to identify resident pathogen strains that could be present in the manufacturing environment and linking that information to potential threats coming in through the supplier network.”
The company offers a family of tests developed on the NGS platform: an authenticity test for food fraud, a universal genetically modified organism (GMO) test and WGS for pathogen strain identification and microbiome. The majority of the testing and analytics work done with manufacturers is currently around safety applications, says Ghorashi, including validating the integrity of the raw ingredients coming into a facility and risk assessment modeling.
Some manufacturers are also using this technology to develop new products. With concerns over products having a longer shelf life—a desire for both retailers and processors—companies are looking at microbial factors affecting spoilage. End users can use Clear Labs WGS test to monitor emerging pathogens.
Clear Labs also offers an extensive molecular food database with more than 2 million sequences and growing.
“We’ve been working on building out our database,” Ghorashi says. “This powers all of our analytics on top of the testing. The beauty of next-gen sequencing is that it generates tons of data—a goldmine for doing things like risk assessment and predictive analytics for the threat of a safety outbreak or pathogen presence.”
3M’s Robayo agrees that WGS has been used effectively for tracking foodborne outbreaks and defining pathogen sources throughout the food chain. It can link a specific pathogen thought to be responsible for an outbreak directly to the food source and identify the pathogen’s species and strain level using a molecular signature. (See “Rapid detection of foodborne bacterial outbreaks” sidebar.) However, she says the data analysis and interpretation still pose a challenge.
“WGS involves an enormous amount of data, and unless there can be significant economies of scale achieved, it’s expensive,” she says. “Its adoption for routine food pathogen testing still needs more development to streamline the analysis and interpretation of data and significantly reduce the costs to price points that are manageable for the industry.”
The benefits of rapid detection methods
Whatever your choice of food safety testing methods, making pathogen protection more automated can have a number of benefits. Automation helps simplify the testing process, which not only saves time, but reduces user error.
“Less human interaction and less need for repeat tests mean lab technicians can spend more time away from the bench and instead concentrate their efforts and attentions on other priorities, allowing for more efficiency in labs and reducing overall costs and time,” says Robayo.
The 3M Molecular Detection System, for instance, utilizes the same assay protocol for all of the pathogens it tests for and accommodates batch processing of up to 96 different tests, per instrument, at the same time. Users need an enriched sample, standard lab equipment, a computer, an assay test and the 3M Molecular Detection instrument. Four instruments can be connected to the same computer, allowing for a total throughput of 384 sample tests to run concurrently.
Once everything is in place, users can run and read tests in approximately 30 minutes with two transfer steps: transferring enriched samples to a prefilled lysis tube rack and placing it on heating and cooling block; and then transferring the lysed samples to color-coded special reagent tubes that are set inside the instrument. After those steps are completed, the automation begins, and results can show up in as little as 15 minutes.
To help the end user and make the pathogen detection process as easy as possible, PathSensors includes proprietary software preloaded with all of its platforms.
“Not only does the software walk the user step by step through the process, it also uses our assay-specific algorithms to interpret test data and provide a positive or negative result,” Fritzinger says.
The software included in many rapid testing systems provides automatic interpretation and analysis of the testing data. In traditional methods, trained technicians are typically needed to perform, interpret and manually record the results.
“People can make mistakes, and multiple technicians might document things differently,” says Robayo. “So, by minimizing the opportunities for technicians to influence results and by capturing data in a uniform manner, food brands can have greater confidence in the fidelity and traceability of the test.”
And when just one mistake can produce such damaging and disastrous effects for food and beverage companies, this can be the difference between success and failure.
Sidebar: Rapid detection of bacterial outbreaks
Researchers from the Georgia Institute of Technology and the US Centers for Disease Control report that a new testing methodology based on metagenomics may speed up identifying microbial causes of foodborne bacterial outbreaks. Metagenomics relies on DNA sequencing and bioinformatics analysis of the resulting sequencing data.
For 20 years, the CDC’s PulseNet, a national laboratory network connecting foodborne illness cases to detect outbreaks, has been using pulsed field gel electrophoresis (PFGE), which requires a bacterial isolate from a patient, animal, food or environmental sample.
“Metagenomics is a way to sequence some or all the bacteria in a patient’s sample without pulling out the bacterial isolate from a sample,” says Dr. Andrew Huang, a microbiologist in CDC’s Enteric Diseases Laboratory Branch. “This method offers an opportunity to save time and go directly to a highly detailed DNA fingerprint for each pathogen.”
In a study comparing the new methodology against traditional culture-based methods with samples from two severe outbreaks of Salmonella, the metagenomics approach identified the correct bacterial culprit and also found a possible coinfection with a second pathogen, Staphylococcus.
“Culture-based approaches are not fast enough and may miss new pathogens that are not represented by available cultures and infections caused by more than one pathogen,” says Kostas Konstantinidis, the Carlton Wilder associate professor at the Georgia Institute of Technology. “When we receive the samples, we can move directly to characterizing them by sequencing the genomes of the microbes that are in the samples without waiting for conventional culturing techniques.”
In the future, according to the researchers, metagenomics technology could improve real-time disease detection and monitoring and provide a better picture of pathogen abundance.
For more information:
Adriana Robayo, 3M, 800-328-1671,
amrobayo@mmm.com, www.3M.com/foodsafety
Jack Fritzinger, PathSensors, 443-557-6150,
info@pathsensors.com, www.pathsensors.com
Mahni Ghorashi, Clear Labs,
inquiries@clearlabs.com, www.clearlabs.com
Michael Koeris, Sample6,
info@sample6.com, www.sample6.com
Looking for a reprint of this article?
From high-res PDFs to custom plaques, order your copy today!





