Food Safety
Automated microbiological testing helps speed up determining if food is safe
The adoption of automated or rapid methods for pathogen testing should include some kind of overall assessment of potential benefits and risks.
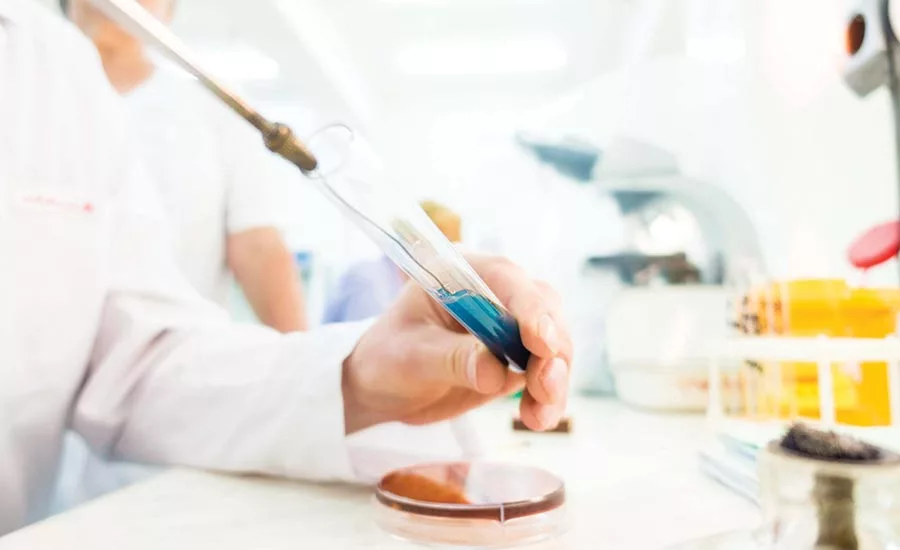

When it comes to pathogen testing, processors must choose from a number of processes, as well as whether the work will be done in-house or through a contract laboratory.
Photo courtesy of Getty Images/Luchschen

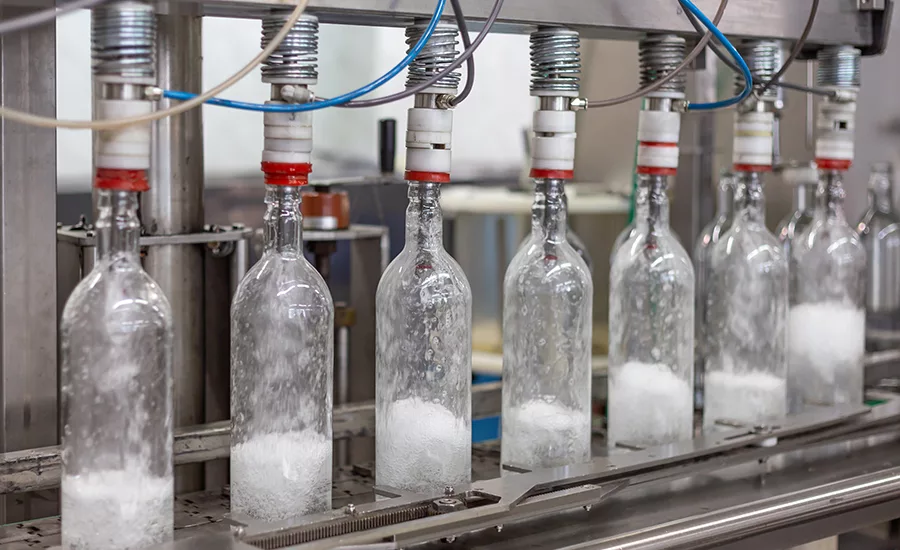
Testing samples of raw or finished products for pathogens presents processors with a number of challenges, including the need to ensure that contract laboratories have equipment properly designed and calibrated to do the specific tests needed.
Photo courtesy of Getty Images/AlexRaths
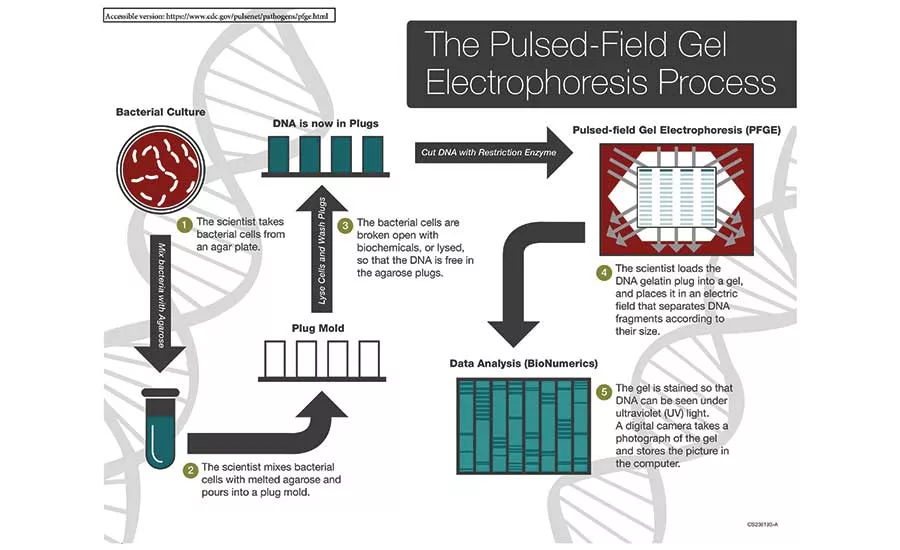
A diagram of the Pulsed-Field Gel Electrophoresis process.
Source: Centers for Disease Control and Prevention
Q&A: LexaGene CEO Dr. Jack Regan
 To meet the increasing needs of consumers and retailers demanding safe food, processors must have a fast and reliable food safety testing and pathogen control program. In this Q&A, Dr. Jack Regan, CEO, LexaGene, shares insights on the new technology available to help automate food safety testing and pathogen control.
To meet the increasing needs of consumers and retailers demanding safe food, processors must have a fast and reliable food safety testing and pathogen control program. In this Q&A, Dr. Jack Regan, CEO, LexaGene, shares insights on the new technology available to help automate food safety testing and pathogen control.
Food Engineering (FE): For food manufacturers, what is the current state of food-borne pathogen detection?
Regan: Food manufacturers test food items as well as have environmental monitoring programs in place to ensure their facilities are clean. For example, when it comes to food items such as ground beef, 25 grams of meat are placed in a bag with enrichment media and are allowed to incubate overnight. For environmental sampling, food-contact surfaces on processing equipment, and non-food contact surfaces (such as floors and drains) are swabbed. The swab is then placed in an enrichment broth. This testing is part of a plant’s overall Hazard Analysis and Critical Control Point (HACCP) plan.
However, for many samples, a “zero tolerance” requirement is imposed on key human pathogens such as Listeria monocytogenes and E. coli—which generally requires an additional culture step to maximize sensitivity, resulting in a longer turnaround time. Following the culture step, a portion of the sample is removed and tested by conventional plate-based technologies or, more commonly, by rapid tests, including immunoassays and PCR-based tests.
FE: What are the drawbacks of using culture-based methods?
Regan: Those methods are very slow. Companies are looking to shorten the enrichment period, from overnight to a few hours to skipping culture entirely, if possible. When you shorten the enrichment period, there are less bacteria present, so immunoassays become less valuable, since they lack the sensitivity of PCR-based assays. However, PCR-based assays are extremely sensitive, as they amplify a single copy of DNA about 1 billion-fold. In addition, PCR does not require a viable pathogen, which is viewed by many as a positive characteristic—since samples that are found to have the DNA from a pathogenic organism are considered ‘high’ risk, and should be treated more cautiously (e.g. a full culture workup). On the other hand, samples with no DNA from a pathogenic organism are considered low risk. PCR also has the advantage that it can be easily multiplexed without harming the sensitivity.
FE: What are some areas in testing methods that are speeding up pathogen detection (in assay preparation, enrichment time, analysis, etc.)?
Regan: The most effort is placed on PCR-based testing, since PCR is so sensitive and amplifies a single copy about 1 billion fold for easy optical detection. Also, there are genomic targets in bacteria that are present at more than 1 copy per cell. So, if bacteria are present—regardless of viability—targeting the mRNA of a highly transcribed gene is more likely to result in a positive test than targeting the bacterium’s gDNA.
FE: How can automated pathogen protection help processors (lower costs, more efficiency, less room for human error, etc.)?
Regan: Automation removes the costs associated with paying an individual to manually process the sample. It also removes some of the variability seen when different microbiologists process the same sample, as many have small variations in technique that can quite drastically change the result. In addition, manual testing is more likely to be prone to false-positives due to user-caused cross-contamination. Automated instruments can run 24/7, which is very helpful for plants that are operational 24 hours a day.
FE: What are some of the potential drawbacks of automated food pathogen detection (occurrence of false positives, need to target a specific pathogen, costs, etc.)?
Regan: Some view PCR testing as “too sensitive.” PCR testing detects the presence of genomic material from different pathogens—but cannot confirm viability. Some meat producers are worried that testing the surfaces of meat processing equipment might too frequently find the presence of DNA from a pathogenic organism. However, it is possible that the source was from bacteria that were killed during processing by a chemical. So, PCR can potentially result in some false alarms, but many feel that this is a better alternative, rather than failing to detect a contaminated food item and sending it to consumers who fall ill.
FE: What is the future of whole genome sequencing and its impact on food safety testing?
Regan: Whole genome sequencing (WGS) is great for identifying where food outbreaks originated from. WGS is a technology that is used after an outbreak has happened, to identify what company should issue the recall. However, WGS is not good for testing on-site at the plant—as it is too time consuming, expensive and technical. This is why it is better to use PCR, which can be easily automated and is much more affordable, in food plants.
Looking for a reprint of this article?
From high-res PDFs to custom plaques, order your copy today!




